H3K4me3 Antibody, SNAP-Certified™ for CUT&RUN and CUT&Tag
{"url":"https://www.epicypher.com/products/antibodies/h3k4me3-antibody-snap-certified-for-cut-run-and-cut-tag","add_this":[{"service":"facebook","annotation":""},{"service":"email","annotation":""},{"service":"print","annotation":""},{"service":"twitter","annotation":""},{"service":"linkedin","annotation":""}],"gtin":null,"id":1119,"bulk_discount_rates":[],"can_purchase":true,"meta_description":"Rabbit monoclonal histone H3K4me3 antibody rigorously tested for robust and reliable performance in CUT&RUN and CUT&Tag","category":["Antibodies","Epigenetics Kits and Reagents/CUTANA™ ChIC / CUT&RUN Assays","Epigenetics Kits and Reagents/CUTANA™ CUT&Tag Assays","Antibodies/CUTANA™ CUT&RUN Compatible Antibodies","Antibodies/CUTANA™ CUT&RUN Antibodies","Antibodies/CUTANA™ CUT&RUN Antibodies/CUTANA™ CUT&RUN Antibodies to Histone PTMs"],"AddThisServiceButtonMeta":"","main_image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/1119/1196/snap-chip-ab__70507.1557259520.1280.1280__90586.1575483123.1280.1280__88802__68533.1701245530.png?c=2","alt":"H3K4me3 Antibody, SNAP-Certified™ for CUT&RUN and CUT&Tag"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=1119","shipping":{"calculated":true},"num_reviews":0,"weight":"0.01 LBS","custom_fields":[{"id":"1288","name":"Pack Size","value":"100 µg"}],"sku":"13-0060","description":"<div class=\"product-general-info\">\n <ul class=\"product-general-info__list-left\">\n <li class=\"product-general-info__list-item\">\n <strong>Type: </strong>Monoclonal [2909-3D7]\n </li>\n <li class=\"product-general-info__list-item\">\n <strong>Host: </strong>Rabbit\n </li>\n <li class=\"product-general-info__list-item\">\n <strong>Applications: </strong>CUT&RUN, CUT&Tag\n </li>\n </ul>\n <ul class=\"product-general-info__list-right\">\n <li class=\"product-general-info__list-item\">\n <strong>Reactivity: </strong>Human, Wide Range\n (Predicted)\n </li>\n <li class=\"product-general-info__list-item\">\n <strong>Format: </strong>Protein A affinity-purified\n </li>\n <li class=\"product-general-info__list-item\">\n <strong>Target Size: </strong>15 kDa\n </li>\n </ul>\n</div>\n\n<div class=\"service_accordion product-droppdown\">\n <div class=\"container\">\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel current\">\n <h3 class=\"sub-title1\">Description</h3>\n <div class=\"ProductDescriptionContainer product-droppdown__section-description-specific\">\n <p>\n This H3K4me3 (histone H3 trimethylated at lysine 4) antibody meets EpiCypher’s lot-specific SNAP-Certified™\n criteria for specificity and efficient target enrichment in both CUT&RUN and CUT&Tag applications. This\n requires <20% cross-reactivity to related histone PTMs determined using the SNAP-CUTANA™ K-MetStat Panel of\n spike-in controls (EpiCypher <a href=\"/products/nucleosomes/snap-cutana-k-metstat-panel\" target=\"_blank\">\n 19-1002</a>, <strong>Figures 1 and 5</strong>). High target efficiency is confirmed by consistent\n genomic enrichment at varying cell inputs: 500k and 50k cells in CUT&RUN (<strong>Figures 2-3</strong>); 100k and 10k cells\n in CUT&Tag (<strong>Figures 6-7</strong>). High efficiency antibodies display similar peak structures (<strong>Figures 3 and 7</strong>) and\n highly conserved genome-wide signal (<strong>Figures 2 and 6</strong>) even at reduced cell numbers. This antibody targets\n histone H3K4me3, which is enriched at active promoters near transcription start sites (TSS) and promotes\n gene activation.\n </p>\n </div>\n </div>\n </div>\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel current\">\n <h3 class=\"sub-title1\">Validation Data - CUT&RUN</h3>\n <div class=\"ProductDescriptionContainer product-droppdown__section-description-specific\">\n <section class=\"image-picker\">\n <div class=\"image-picker__left\">\n <div class=\"image-picker__main-content_active image-picker__main-content\">\n <div class=\"image-picker__header-content\">\n <button class=\"image-picker__left-arrow\">\n <svg class=\"image-picker__svg-left\" width=\"24\" height=\"24\" viewBox=\"0 0 24 24\">\n <path d=\"M16.67 0l2.83 2.829-9.339 9.175 9.339 9.167-2.83 2.829-12.17-11.996z\" />\n </svg>\n </button>\n <a href=\"/content/images/products/antibodies/13-0060_snap_specificity_cr.jpeg\"\n target=\"_blank\" class=\"image-picker__main-image-link\"><img loading=\"lazy\"\n alt=\"13-0060_snap_specificity_cr\"\n src=\"/content/images/products/antibodies/13-0060_snap_specificity_cr.jpeg\"\n class=\"image-picker__main-image\" />\n <span class=\"image-picker__main-image-caption\">(Click to enlarge)</span></a>\n <button class=\"image-picker__right-arrow\">\n <svg class=\"image-picker__svg-right\" width=\"24\" height=\"24\" viewBox=\"0 0 24 24\">\n <path d=\"M7.33 24l-2.83-2.829 9.339-9.175-9.339-9.167 2.83-2.829 12.17 11.996z\" />\n </svg>\n </button>\n </div>\n <p>\n <span class=\"image-picker__span-content\"><strong>Figure 1: SNAP specificity analysis in\n CUT&RUN</strong><br />\n CUT&RUN was performed as described in\n <strong>Figure 4</strong>. CUT&RUN sequencing reads\n were aligned to the unique DNA barcodes corresponding\n to each nucleosome in the K-MetStat panel (x-axis).\n Data are expressed as a percent relative to on-target\n recovery (H3K4me3 set to 100%). The antibody showed highly specific recovery of H3K4me3 spike-in\n nucleosomes at both 500k and 50k cells.\n </span>\n </p>\n </div>\n <div class=\"image-picker__main-content\">\n <div class=\"image-picker__header-content\">\n <button class=\"image-picker__left-arrow\">\n <svg class=\"image-picker__svg-left\" width=\"24\" height=\"24\" viewBox=\"0 0 24 24\">\n <path d=\"M16.67 0l2.83 2.829-9.339 9.175 9.339 9.167-2.83 2.829-12.17-11.996z\" />\n </svg>\n </button>\n <a href=\"/content/images/products/antibodies/13-0060_genome-wide_enrichment_cr.jpeg\"\n target=\"_blank\" class=\"image-picker__main-image-link\"><img loading=\"lazy\"\n alt=\"13-0060_genome-wide_enrichment_cr\"\n src=\"/content/images/products/antibodies/13-0060_genome-wide_enrichment_cr.jpeg\"\n class=\"image-picker__main-image\" />\n <span class=\"image-picker__main-image-caption\">(Click to enlarge)</span></a>\n <button class=\"image-picker__right-arrow\">\n <svg class=\"image-picker__svg-right\" width=\"24\" height=\"24\" viewBox=\"0 0 24 24\">\n <path d=\"M7.33 24l-2.83-2.829 9.339-9.175-9.339-9.167 2.83-2.829 12.17 11.996z\" />\n </svg>\n </button>\n </div>\n <p>\n <span class=\"image-picker__span-content\"><strong>Figure 2: CUT&RUN genome-wide\n enrichment</strong><br />\n CUT&RUN was performed as described in\n <strong>Figure 4</strong>. Sequence reads were aligned\n to 18,793 annotated transcription start sites (TSSs ±\n 2 kbp). Signal enrichment was sorted from highest to\n lowest (top to bottom) relative to the H3K4me3 - 500k\n cells reaction (all gene rows aligned). High, medium,\n and low intensity are shown in red, yellow, and blue,\n respectively. H3K4me3 antibodies produced the expected\n enrichment pattern, which was consistent between 500k\n and 50k cells and greater than the IgG negative\n control.\n </span>\n </p>\n </div>\n <div class=\"image-picker__main-content\">\n <div class=\"image-picker__header-content\">\n <button class=\"image-picker__left-arrow\">\n <svg class=\"image-picker__svg-left\" width=\"24\" height=\"24\" viewBox=\"0 0 24 24\">\n <path d=\"M16.67 0l2.83 2.829-9.339 9.175 9.339 9.167-2.83 2.829-12.17-11.996z\" />\n </svg>\n </button>\n <a href=\"/content/images/products/antibodies/13-0060_browser_tracks_cr.jpeg\"\n target=\"_blank\" class=\"image-picker__main-image-link\">\n <img loading=\"lazy\" alt=\"13-0060_browser_tracks_cr\"\n src=\"/content/images/products/antibodies/13-0060_browser_tracks_cr.jpeg\"\n class=\"image-picker__main-image\" />\n <span class=\"image-picker__main-image-caption\">(Click to enlarge)</span>\n </a>\n <button class=\"image-picker__right-arrow\">\n <svg class=\"image-picker__svg-right\" width=\"24\" height=\"24\" viewBox=\"0 0 24 24\">\n <path d=\"M7.33 24l-2.83-2.829 9.339-9.175-9.339-9.167 2.83-2.829 12.17 11.996z\" />\n </svg>\n </button>\n </div>\n <p>\n <span class=\"image-picker__span-content\">\n <strong>Figure 3: H3K4me3 CUT&RUN representative browser tracks</strong><br />\n CUT&RUN was performed as described in\n <strong>Figure 4</strong>. Gene browser shots were\n generated using the Integrative Genomics Viewer (IGV,\n Broad Institute). H3K4me3 antibody tracks display sharp peaks at gene promoters, consistent with the\n biological function of this PTM. Similar results in peak structure and\n location were observed for both 500k and 50k cell\n inputs.\n </span>\n </p>\n </div>\n <div class=\"image-picker__main-content\">\n <div class=\"image-picker__header-content\">\n <button class=\"image-picker__left-arrow\">\n <svg class=\"image-picker__svg-left\" width=\"24\" height=\"24\" viewBox=\"0 0 24 24\">\n <path d=\"M16.67 0l2.83 2.829-9.339 9.175 9.339 9.167-2.83 2.829-12.17-11.996z\" />\n </svg>\n </button>\n <a href=\"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/original/image-manager/13-0060-figure4.png?t=1700792532\"\n target=\"_blank\" class=\"image-picker__main-image-link\">\n <img loading=\"lazy\" alt=\"cut-and-run-methods\"\n src=\"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/original/image-manager/13-0060-figure4.png?t=1700792532\"\n class=\"image-picker__main-image\" />\n <span class=\"image-picker__main-image-caption\">(Click to enlarge)</span>\n </a>\n <button class=\"image-picker__right-arrow\">\n <svg class=\"image-picker__svg-right\" width=\"24\" height=\"24\" viewBox=\"0 0 24 24\">\n <path d=\"M7.33 24l-2.83-2.829 9.339-9.175-9.339-9.167 2.83-2.829 12.17 11.996z\" />\n </svg>\n </button>\n </div>\n <p>\n <span class=\"image-picker__span-content\">\n <strong>Figure 4: CUT&RUN methods</strong><br />\n CUT&RUN was performed on 500k and 50k K562 cells with\n the SNAP-CUTANA™ K-MetStat Panel (EpiCypher\n <a href=\"https://www.epicypher.com/products/nucleosomes/snap-cutana-k-metstat-panel\"\n target=\"_blank\">19-1002</a>) spiked-in prior to the addition of 0.5 µg of either\n IgG negative control (EpiCypher\n <a href=\"https://www.epicypher.com/products/nucleosomes/snap-cutana-spike-in-controls/cutana-rabbit-igg-cut-run-negative-control-antibody\"\n target=\"_blank\">13-0042</a>) or H3K4me3 antibodies. The experiment was performed\n using the CUTANA™ ChIC/CUT&RUN Kit v3 (EpiCypher\n <a href=\"https://www.epicypher.com/products/epigenetics-reagents-and-assays/cutana-chic-cut-and-run-kit\"\n target=\"_blank\">14-1048</a>). Library preparation was performed with 5 ng of\n CUT&RUN enriched DNA (or the total amount recovered if\n less than 5 ng) using the CUTANA™ CUT&RUN Library Prep\n Kit (EpiCypher\n <a href=\"https://www.epicypher.com/products/epigenetics-reagents-and-assays/cutana-cut-and-run-library-prep-kit\"\n target=\"_blank\">14-1001/14-1002</a>). Both kit protocols were adapted for high\n throughput Tecan liquid handling. Libraries were run\n on an Illumina NextSeq2000 with paired-end sequencing\n (2x50 bp). Sample sequencing depth was 3.5 million reads (IgG 500k cell input), 4.0 million reads\n (IgG 50k cell input), 5.6 million reads (H3K4me3 500k cell input), and 2.8 million reads (H3K4me3\n 50k cell input). Data were\n aligned to the hg19 genome using Bowtie2. Data were\n filtered to remove duplicates, multi-aligned reads,\n and ENCODE DAC Exclusion List regions.\n </span>\n </p>\n </div>\n </div>\n <aside class=\"image-picker__right\">\n <div class=\"image-picker__gallery\">\n <img loading=\"lazy\" alt=\"13-0055-specificity-analysis\"\n src=\"/content/images/products/antibodies/13-0060_snap_specificity_cr.jpeg\" width=\"200\"\n class=\"image-picker__side-image image-picker__side-image_active\" role=\"button\" />\n <img loading=\"lazy\" alt=\"13-0055-genome-wide-enrichment\"\n src=\"/content/images/products/antibodies/13-0060_genome-wide_enrichment_cr.jpeg\"\n class=\"image-picker__side-image\" role=\"button\" />\n <img loading=\"lazy\" alt=\"13-0055-representative-browser-tracks\"\n src=\"/content/images/products/antibodies/13-0060_browser_tracks_cr.jpeg\"\n class=\"image-picker__side-image\" role=\"button\" />\n <img loading=\"lazy\" alt=\"cut-and-run-methods\"\n src=\"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/original/image-manager/13-0060-figure4.png?t=1700792532\"\n class=\"image-picker__side-image\" role=\"button\" />\n </div>\n </aside>\n </section>\n </div>\n </div>\n </div>\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel current\">\n <h3 class=\"sub-title1\">Validation Data - CUT&Tag</h3>\n <div class=\"ProductDescriptionContainer product-droppdown__section-description-specific\">\n <section class=\"image-picker\">\n <div class=\"image-picker__left\">\n <div class=\"image-picker__main-content_active image-picker__main-content\">\n <div class=\"image-picker__header-content\">\n <button class=\"image-picker__left-arrow\">\n <svg class=\"image-picker__svg-left\" width=\"24\" height=\"24\" viewBox=\"0 0 24 24\">\n <path d=\"M16.67 0l2.83 2.829-9.339 9.175 9.339 9.167-2.83 2.829-12.17-11.996z\" />\n </svg>\n </button>\n <a href=\"/content/images/products/antibodies/13-0060_snap_specificity_ct.jpeg\"\n target=\"_blank\" class=\"image-picker__main-image-link\">\n <img loading=\"lazy\" alt=\"13-0060_snap_specificity_ct\"\n src=\"/content/images/products/antibodies/13-0060_snap_specificity_ct.jpeg\"\n class=\"image-picker__main-image\" />\n <span class=\"image-picker__main-image-caption\">(Click to enlarge)</span>\n </a>\n <button class=\"image-picker__right-arrow\">\n <svg class=\"image-picker__svg-right\" width=\"24\" height=\"24\" viewBox=\"0 0 24 24\">\n <path d=\"M7.33 24l-2.83-2.829 9.339-9.175-9.339-9.167 2.83-2.829 12.17 11.996z\" />\n </svg>\n </button>\n </div>\n <p>\n <span class=\"image-picker__span-content\">\n <strong>Figure 5: SNAP specificity analysis in\n CUT&Tag</strong><br />\n CUT&Tag was performed as described in\n <strong>Figure 8</strong>. CUT&Tag sequencing reads\n were aligned to the unique DNA barcodes corresponding\n to each nucleosome in the K-MetStat panel (x-axis).\n Data are expressed as a percent relative to on-target\n recovery (H3K4me3 set to 100%). The antibody showed highly specific recovery of H3K4me3 spike-in\n nucleosomes at both 100k and 10k nuclei.\n </span>\n </p>\n </div>\n <div class=\"image-picker__main-content\">\n <div class=\"image-picker__header-content\">\n <button class=\"image-picker__left-arrow\">\n <svg class=\"image-picker__svg-left\" width=\"24\" height=\"24\" viewBox=\"0 0 24 24\">\n <path d=\"M16.67 0l2.83 2.829-9.339 9.175 9.339 9.167-2.83 2.829-12.17-11.996z\" />\n </svg>\n </button>\n <a href=\"/content/images/products/antibodies/13-0060_genome-wide_enrichment_ct.jpeg\"\n target=\"_blank\" class=\"image-picker__main-image-link\">\n <img loading=\"lazy\" alt=\"13-0060_genome-wide_enrichment_ct\"\n src=\"/content/images/products/antibodies/13-0060_genome-wide_enrichment_ct.jpeg\"\n class=\"image-picker__main-image\" />\n <span class=\"image-picker__main-image-caption\">(Click to enlarge)</span>\n </a>\n <button class=\"image-picker__right-arrow\">\n <svg class=\"image-picker__svg-right\" width=\"24\" height=\"24\" viewBox=\"0 0 24 24\">\n <path d=\"M7.33 24l-2.83-2.829 9.339-9.175-9.339-9.167 2.83-2.829 12.17 11.996z\" />\n </svg>\n </button>\n </div>\n <p>\n <span class=\"image-picker__span-content\">\n <strong>Figure 6: CUT&Tag genome-wide enrichment </strong><br />\n CUT&Tag was performed as described in\n <strong>Figure 8</strong>. Sequence reads were aligned\n to 18,793 annotated transcription start sites (TSSs ±\n 2 kbp). Signal enrichment was sorted from highest to\n lowest (top to bottom) relative to the H3K4me3 - 100k\n nuclei reaction (all gene rows aligned). High, medium,\n and low intensity are shown in red, yellow, and blue,\n respectively. H3K4me3 antibodies produced the expected\n enrichment pattern, which was consistent between 100k\n and 10k nuclei and greater than the IgG negative\n control.\n </span>\n </p>\n </div>\n <div class=\"image-picker__main-content\">\n <div class=\"image-picker__header-content\">\n <button class=\"image-picker__left-arrow\">\n <svg class=\"image-picker__svg-left\" width=\"24\" height=\"24\" viewBox=\"0 0 24 24\">\n <path d=\"M16.67 0l2.83 2.829-9.339 9.175 9.339 9.167-2.83 2.829-12.17-11.996z\" />\n </svg>\n </button>\n <a href=\"/content/images/products/antibodies/13-0060_browser_tracks_ct.jpeg\"\n target=\"_blank\" class=\"image-picker__main-image-link\">\n <img loading=\"lazy\" alt=\"13-0060_browser_tracks_ct\"\n src=\"/content/images/products/antibodies/13-0060_browser_tracks_ct.jpeg\"\n class=\"image-picker__main-image\" />\n <span class=\"image-picker__main-image-caption\">(Click to enlarge)</span>\n </a>\n <button class=\"image-picker__right-arrow\">\n <svg class=\"image-picker__svg-right\" width=\"24\" height=\"24\" viewBox=\"0 0 24 24\">\n <path d=\"M7.33 24l-2.83-2.829 9.339-9.175-9.339-9.167 2.83-2.829 12.17 11.996z\" />\n </svg>\n </button>\n </div>\n <p>\n <span class=\"image-picker__span-content\">\n <strong>Figure 7: H3K4me3 CUT&Tag representative browser tracks</strong><br />\n CUT&Tag was performed as described in\n <strong>Figure 8</strong>. Gene browser shots were\n generated using the Integrative Genomics Viewer (IGV,\n Broad Institute). H3K4me3 antibody tracks display sharp peaks at gene promoters, consistent with the\n biological function of this PTM. Similar results in peak structure and\n location were observed for both 100k and 10k nuclei\n inputs.\n </span>\n </p>\n </div>\n <div class=\"image-picker__main-content\">\n <div class=\"image-picker__header-content\">\n <button class=\"image-picker__left-arrow\">\n <svg class=\"image-picker__svg-left\" width=\"24\" height=\"24\" viewBox=\"0 0 24 24\">\n <path d=\"M16.67 0l2.83 2.829-9.339 9.175 9.339 9.167-2.83 2.829-12.17-11.996z\" />\n </svg>\n </button>\n <a href=\"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/original/image-manager/13-0060-figure8.png?t=1700792533\"\n target=\"_blank\" class=\"image-picker__main-image-link\">\n <img loading=\"lazy\" alt=\"cut-and-tag-methods\"\n src=\"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/original/image-manager/13-0060-figure8.png?t=1700792533\"\n class=\"image-picker__main-image\" />\n <span class=\"image-picker__main-image-caption\">(Click to enlarge)</span>\n </a>\n <button class=\"image-picker__right-arrow\">\n <svg class=\"image-picker__svg-right\" width=\"24\" height=\"24\" viewBox=\"0 0 24 24\">\n <path d=\"M7.33 24l-2.83-2.829 9.339-9.175-9.339-9.167 2.83-2.829 12.17 11.996z\" />\n </svg>\n </button>\n </div>\n <p>\n <span class=\"image-picker__span-content\">\n <strong>Figure 8: CUT&Tag methods</strong><br />\n CUT&Tag was performed on 100k and 10k K562 nuclei with\n the SNAP-CUTANA™ K-MetStat Panel (EpiCypher\n <a href=\"https://www.epicypher.com/products/nucleosomes/snap-cutana-k-metstat-panel\"\n target=\"_blank\">19-1002</a>) spiked-in prior to the addition of 0.5 µg of either\n IgG negative control (EpiCypher\n <a href=\"https://www.epicypher.com/products/nucleosomes/snap-cutana-spike-in-controls/cutana-rabbit-igg-cut-run-negative-control-antibody\"\n target=\"_blank\">13-0042</a>) or H3K4me3 antibodies. The experiment was performed\n using the CUTANA™ CUT&Tag Kit v1 (EpiCypher\n <a href=\"https://www.epicypher.com/products/epigenetics-kits-and-reagents/cutana-cut-and-tag-kit\"\n target=\"_blank\">14-1102/14-1103</a>). Libraries were run on an Illumina NextSeq2000 with\n paired-end sequencing (2x50 bp). Sample sequencing depth was 1.3 million reads (IgG 100k nuclei\n input), 2.0 million reads (IgG 10k nuclei input), 6.9 million reads (H3K4me3 100k nuclei input), and\n 10.6 million reads (H3K4me3 10k nuclei input). Data were aligned to the\n hg19 genome using Bowtie2. Data were filtered to\n remove duplicates, multi-aligned reads, and ENCODE DAC\n Exclusion List regions.\n </span>\n </p>\n </div>\n </div>\n <aside class=\"image-picker__right\">\n <div class=\"image-picker__gallery\">\n <img loading=\"lazy\" alt=\"13-0060_snap_specificity_ct\"\n src=\"/content/images/products/antibodies/13-0060_snap_specificity_ct.jpeg\"\n class=\"image-picker__side-image\" role=\"button\" />\n <img loading=\"lazy\" alt=\"13-0060_genome-wide_enrichment_ct\"\n src=\"/content/images/products/antibodies/13-0060_genome-wide_enrichment_ct.jpeg\"\n class=\"image-picker__side-image\" role=\"button\" />\n <img loading=\"lazy\" alt=\"13-0060_browser_tracks_ct\"\n src=\"/content/images/products/antibodies/13-0060_browser_tracks_ct.jpeg\"\n class=\"image-picker__side-image\" role=\"button\" />\n <img loading=\"lazy\" alt=\"cut-and-tag-methods\"\n src=\"/content/images/products/methods/cut-and-tag-methods.png\" class=\"image-picker__side-image\"\n role=\"button\" />\n </div>\n </aside>\n </section>\n </div>\n </div>\n </div>\n\n\n\n\n\n\n\n <!-- <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel\">\n <h3 class=\"sub-title1\">Validation Data - CUT&Tag</h3>\n <div class=\"ProductDescriptionContainer product-droppdown__section-description-specific\" style=\"display: none\">\n <p></p>\n <section class=\"image-picker\">\n <div class=\"image-picker__left\">\n <div class=\"image-picker__main-content_active image-picker__main-content\">\n <div class=\"image-picker__header-content\">\n <button class=\"image-picker__left-arrow\">\n <svg class=\"image-picker__svg-left\" width=\"24\" height=\"24\" viewBox=\"0 0 24 24\">\n <path d=\"M16.67 0l2.83 2.829-9.339 9.175 9.339 9.167-2.83 2.829-12.17-11.996z\" />\n </svg>\n </button>\n <a href=\"/content/images/products/antibodies/13-0060_snap_specificity_ct.jpeg\"\n target=\"_blank\" class=\"image-picker__main-image-link\">\n <img loading=\"lazy\" alt=\"13-0060_snap_specificity_ct\"\n src=\"/content/images/products/antibodies/13-0060_snap_specificity_ct.jpeg\"\n class=\"image-picker__main-image\" />\n <span class=\"image-picker__main-image-caption\">(Click to enlarge)</span>\n </a>\n <button class=\"image-picker__right-arrow\">\n <svg class=\"image-picker__svg-right\" width=\"24\" height=\"24\" viewBox=\"0 0 24 24\">\n <path d=\"M7.33 24l-2.83-2.829 9.339-9.175-9.339-9.167 2.83-2.829 12.17 11.996z\" />\n </svg>\n </button>\n </div>\n <p>\n <span class=\"image-picker__span-content\">\n <strong>Figure 5: SNAP specificity analysis in\n CUT&Tag</strong><br />\n CUT&Tag was performed as described in\n <strong>Figure 8</strong>. CUT&Tag sequencing reads\n were aligned to the unique DNA barcodes corresponding\n to each nucleosome in the K-MetStat panel (x-axis).\n Data are expressed as a percent relative to on-target\n recovery (H3K4me3 set to 100%). The antibody showed highly specific recovery of H3K4me3 spike-in\n nucleosomes at both 100k and 10k nuclei.\n </span>\n </p>\n </div>\n <div class=\"image-picker__main-content\">\n <div class=\"image-picker__header-content\">\n <button class=\"image-picker__left-arrow\">\n <svg class=\"image-picker__svg-left\" width=\"24\" height=\"24\" viewBox=\"0 0 24 24\">\n <path d=\"M16.67 0l2.83 2.829-9.339 9.175 9.339 9.167-2.83 2.829-12.17-11.996z\" />\n </svg>\n </button>\n <a href=\"/content/images/products/antibodies/13-0060_genome-wide_enrichment_ct.jpeg\"\n target=\"_blank\" class=\"image-picker__main-image-link\">\n <img loading=\"lazy\" alt=\"13-0060_genome-wide_enrichment_ct\"\n src=\"/content/images/products/antibodies/13-0060_genome-wide_enrichment_ct.jpeg\"\n class=\"image-picker__main-image\" />\n <span class=\"image-picker__main-image-caption\">(Click to enlarge)</span>\n </a>\n <button class=\"image-picker__right-arrow\">\n <svg class=\"image-picker__svg-right\" width=\"24\" height=\"24\" viewBox=\"0 0 24 24\">\n <path d=\"M7.33 24l-2.83-2.829 9.339-9.175-9.339-9.167 2.83-2.829 12.17 11.996z\" />\n </svg>\n </button>\n </div>\n <p>\n <span class=\"image-picker__span-content\">\n <strong>Figure 6: CUT&Tag genome-wide enrichment </strong><br />\n CUT&Tag was performed as described in\n <strong>Figure 8</strong>. Sequence reads were aligned\n to 18,793 annotated transcription start sites (TSSs ±\n 2 kbp). Signal enrichment was sorted from highest to\n lowest (top to bottom) relative to the H3K4me3 - 100k\n nuclei reaction (all gene rows aligned). High, medium,\n and low intensity are shown in red, yellow, and blue,\n respectively. H3K4me3 antibodies produced the expected\n enrichment pattern, which was consistent between 100k\n and 10k nuclei and greater than the IgG negative\n control.\n </span>\n </p>\n </div>\n <div class=\"image-picker__main-content\">\n <div class=\"image-picker__header-content\">\n <button class=\"image-picker__left-arrow\">\n <svg class=\"image-picker__svg-left\" width=\"24\" height=\"24\" viewBox=\"0 0 24 24\">\n <path d=\"M16.67 0l2.83 2.829-9.339 9.175 9.339 9.167-2.83 2.829-12.17-11.996z\" />\n </svg>\n </button>\n <a href=\"/content/images/products/antibodies/13-0060_browser_tracks_ct.jpeg\"\n target=\"_blank\" class=\"image-picker__main-image-link\">\n <img loading=\"lazy\" alt=\"13-0060_browser_tracks_ct\"\n src=\"/content/images/products/antibodies/13-0060_browser_tracks_ct.jpeg\"\n class=\"image-picker__main-image\" />\n <span class=\"image-picker__main-image-caption\">(Click to enlarge)</span>\n </a>\n <button class=\"image-picker__right-arrow\">\n <svg class=\"image-picker__svg-right\" width=\"24\" height=\"24\" viewBox=\"0 0 24 24\">\n <path d=\"M7.33 24l-2.83-2.829 9.339-9.175-9.339-9.167 2.83-2.829 12.17 11.996z\" />\n </svg>\n </button>\n </div>\n <p>\n <span class=\"image-picker__span-content\">\n <strong>Figure 7: H3K4me3 CUT&Tag representative browser tracks</strong><br />\n CUT&Tag was performed as described in\n <strong>Figure 8</strong>. Gene browser shots were\n generated using the Integrative Genomics Viewer (IGV,\n Broad Institute). H3K4me3 antibody tracks display sharp peaks at gene promoters, consistent with the\n biological function of this PTM. Similar results in peak structure and\n location were observed for both 100k and 10k nuclei\n inputs.\n </span>\n </p>\n </div>\n <div class=\"image-picker__main-content\">\n <div class=\"image-picker__header-content\">\n <button class=\"image-picker__left-arrow\">\n <svg class=\"image-picker__svg-left\" width=\"24\" height=\"24\" viewBox=\"0 0 24 24\">\n <path d=\"M16.67 0l2.83 2.829-9.339 9.175 9.339 9.167-2.83 2.829-12.17-11.996z\" />\n </svg>\n </button>\n <a href=\"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/original/image-manager/13-0060-figure8.png?t=1700792533\"\n target=\"_blank\" class=\"image-picker__main-image-link\">\n <img loading=\"lazy\" alt=\"cut-and-tag-methods\"\n src=\"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/original/image-manager/13-0060-figure8.png?t=1700792533\"\n class=\"image-picker__main-image\" />\n <span class=\"image-picker__main-image-caption\">(Click to enlarge)</span>\n </a>\n <button class=\"image-picker__right-arrow\">\n <svg class=\"image-picker__svg-right\" width=\"24\" height=\"24\" viewBox=\"0 0 24 24\">\n <path d=\"M7.33 24l-2.83-2.829 9.339-9.175-9.339-9.167 2.83-2.829 12.17 11.996z\" />\n </svg>\n </button>\n </div>\n <p>\n <span class=\"image-picker__span-content\">\n <strong>Figure 8: CUT&Tag methods</strong><br />\n CUT&Tag was performed on 100k and 10k K562 nuclei with\n the SNAP-CUTANA™ K-MetStat Panel (EpiCypher\n <a href=\"https://www.epicypher.com/products/nucleosomes/snap-cutana-k-metstat-panel\"\n target=\"_blank\">19-1002</a>) spiked-in prior to the addition of 0.5 µg of either\n IgG negative control (EpiCypher\n <a href=\"https://www.epicypher.com/products/nucleosomes/snap-cutana-spike-in-controls/cutana-rabbit-igg-cut-run-negative-control-antibody\"\n target=\"_blank\">13-0042</a>) or H3K4me3 antibodies. The experiment was performed\n using the CUTANA™ CUT&Tag Kit v1 (EpiCypher\n <a href=\"https://www.epicypher.com/products/epigenetics-kits-and-reagents/cutana-cut-and-tag-kit\"\n target=\"_blank\">14-1102/14-1103</a>). Libraries were run on an Illumina NextSeq2000 with\n paired-end sequencing (2x50 bp). Sample sequencing depth was 1.3 million reads (IgG 100k nuclei\n input), 2.0 million reads (IgG 10k nuclei input), 6.9 million reads (H3K4me3 100k nuclei input), and\n 10.6 million reads (H3K4me3 10k nuclei input). Data were aligned to the\n hg19 genome using Bowtie2. Data were filtered to\n remove duplicates, multi-aligned reads, and ENCODE DAC\n Exclusion List regions.\n </span>\n </p>\n </div>\n </div>\n <aside class=\"image-picker__right\">\n <div class=\"image-picker__gallery\">\n <img loading=\"lazy\" alt=\"13-0060_snap_specificity_ct\"\n src=\"/content/images/products/antibodies/13-0060_snap_specificity_ct.jpeg\"\n class=\"image-picker__side-image\" role=\"button\" />\n <img loading=\"lazy\" alt=\"13-0060_genome-wide_enrichment_ct\"\n src=\"/content/images/products/antibodies/13-0060_genome-wide_enrichment_ct.jpeg\"\n class=\"image-picker__side-image\" role=\"button\" />\n <img loading=\"lazy\" alt=\"13-0060_browser_tracks_ct\"\n src=\"/content/images/products/antibodies/13-0060_browser_tracks_ct.jpeg\"\n class=\"image-picker__side-image\" role=\"button\" />\n <img loading=\"lazy\" alt=\"cut-and-tag-methods\"\n src=\"/content/images/products/methods/cut-and-tag-methods.png\" class=\"image-picker__side-image\"\n role=\"button\" />\n </div>\n </aside>\n </section>\n </div>\n </div>\n </div> -->\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel\">\n <h3 class=\"sub-title1\">Technical Information</h3>\n <div class=\"ProductDescriptionContainer product-droppdown__section-description\">\n <div class=\"product-tech-info\">\n <div class=\"product-tech-info__line-item\">\n <div class=\"product-tech-info__line-item-left\">\n <b>Immunogen</b>\n </div>\n <div class=\"product-tech-info__line-item-right\">\n A synthetic peptide corresponding to histone H3\n trimethylated at lysine 4\n </div>\n </div>\n <div class=\"product-tech-info__line-item\">\n <div class=\"product-tech-info__line-item-left\">\n <b>Storage</b>\n </div>\n <div class=\"product-tech-info__line-item-right\">\n Stable for 1 year at 4ºC from date of receipt\n </div>\n </div>\n <div class=\"product-tech-info__line-item\">\n <div class=\"product-tech-info__line-item-left\">\n <b>Formulation</b>\n </div>\n <div class=\"product-tech-info__line-item-right\">\n Protein A affinity-purified recombinant monoclonal antibody in borate buffered\n saline pH 8.0, 0.09% sodium azide\n </div>\n </div>\n </div>\n </div>\n </div>\n </div>\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel\">\n <h3 class=\"sub-title1\">Recommended Dilution</h3>\n <div class=\"ProductDescriptionContainer product-droppdown__section-description\">\n <div class=\"product-tech-info\">\n <div class=\"product-tech-info__line-item\">\n <div class=\"product-tech-info__line-item-left\">\n <b>CUT&RUN</b>\n </div>\n <div class=\"product-tech-info__line-item-right\">\n 0.5 µg per reaction\n </div>\n </div>\n <div class=\"product-tech-info__line-item\">\n <div class=\"product-tech-info__line-item-left\">\n <b>CUT&Tag</b>\n </div>\n <div class=\"product-tech-info__line-item-right\">\n 0.5 µg per reaction\n </div>\n </div>\n </div>\n </div>\n </div>\n </div>\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel\">\n <h3 class=\"sub-title1\">Gene & Protein Information</h3>\n <div class=\"ProductDescriptionContainer product-droppdown__section-description\">\n <div class=\"product-tech-info\">\n <div class=\"product-tech-info__line-item\">\n <div class=\"product-tech-info__line-item-left\">\n <b>Uniprot ID</b>\n </div>\n <div class=\"product-tech-info__line-item-right\">\n H3.1 - P68431\n </div>\n </div>\n <div class=\"product-tech-info__line-item\">\n <div class=\"product-tech-info__line-item-left\">\n <b>Alternate Names</b>\n </div>\n <div class=\"product-tech-info__line-item-right\">\n H3, H3/a, H3/b, H3/d\n </div>\n </div>\n </div>\n </div>\n </div>\n </div>\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel\">\n <h3 class=\"sub-title1\">Documents & Resources</h3>\n <div class=\"ProductDescriptionContainer product-droppdown__section-description\">\n <div class=\"product-documents\">\n <a href=\"/content/documents/tds/13-0060.pdf\" target=\"_blank\" class=\"product-documents__link\">\n <svg version=\"1.1\" id=\"Layer_1\" xmlns=\"http://www.w3.org/2000/svg\"\n xmlns:xlink=\"http://www.w3.org/1999/xlink\" x=\"0px\" y=\"0px\" viewBox=\"0 0 228 240\"\n style=\"enable-background: new 0 0 228 240\" xml:space=\"preserve\" class=\"product-documents__icon\"\n alt=\"16-0030 Datasheet\">\n <g>\n <path class=\"product-documents__svg-pdf\"\n d=\"M191.92,68.77l-47.69-47.69c-1.33-1.33-3.12-2.08-5.01-2.08H45.09C41.17,19,38,22.17,38,26.09v184.36\n c0,3.92,3.17,7.09,7.09,7.09h141.82c3.92,0,7.09-3.17,7.09-7.09V73.8C194,71.92,193.25,70.1,191.92,68.77z M177.65,77.06h-41.7\n v-41.7L177.65,77.06z M178.05,201.59H53.95V34.95h66.92v47.86c0,5.14,4.17,9.31,9.31,9.31h47.86V201.59z\" />\n </g>\n <rect x=\"20\" y=\"112\" class=\"product-documents__svg-background\" width=\"146\" height=\"76\" />\n <g>\n <path class=\"product-documents__svg-pdf\" d=\"M23.83,125.68h22.36c5.29,0,9.41,1.33,12.35,4c2.94,2.67,4.42,6.39,4.42,11.18c0,4.78-1.47,8.51-4.42,11.18\n c-2.94,2.67-7.06,4-12.35,4H34.59v18.29H23.83V125.68z M44.81,147.9c5.38,0,8.07-2.32,8.07-6.97c0-2.39-0.67-4.16-2-5.31\n c-1.33-1.15-3.36-1.73-6.07-1.73H34.59v14.01H44.81z\" />\n <path class=\"product-documents__svg-pdf\"\n d=\"M69.92,125.68h18.91c5.29,0,9.84,0.97,13.66,2.9c3.82,1.93,6.74,4.72,8.76,8.35\n c2.02,3.63,3.04,7.98,3.04,13.04c0,5.06-1,9.42-3,13.08c-2,3.66-4.91,6.45-8.73,8.38c-3.82,1.93-8.4,2.9-13.73,2.9H69.92V125.68z\n M88.07,165.63c10.35,0,15.52-5.22,15.52-15.66c0-10.4-5.17-15.59-15.52-15.59h-7.38v31.26H88.07z\" />\n <path class=\"product-documents__svg-pdf\"\n d=\"M122.57,125.68h32.84v8.49h-22.22v11.18h20.84v8.49h-20.84v20.49h-10.63V125.68z\" />\n </g>\n </svg>\n <span class=\"product-documents__info\">Technical Datasheet</span>\n </a>\n </div>\n </div>\n </div>\n </div>\n </div>\n</div>","tags":[],"warranty":"","price":{"without_tax":{"formatted":"$545.00","value":545,"currency":"USD"},"tax_label":"Sales Tax"},"detail_messages":"","availability":"","page_title":"H3K4me3 Antibody | SNAP-Certified™ for CUT&RUN and CUT&Tag","cart_url":"https://www.epicypher.com/cart.php","max_purchase_quantity":0,"mpn":null,"upc":null,"options":[],"related_products":[{"id":694,"sku":null,"name":"CUTANA™ pAG-MNase for ChIC/CUT&RUN Workflows","url":"https://www.epicypher.com/products/epigenetics-reagents-and-assays/cutana-pag-mnase-for-chic-cut-and-run-workflows","availability":"","rating":null,"brand":{"name":null},"category":["Epigenetics Kits and Reagents","Epigenetics Kits and Reagents/CUTANA™ ChIC / CUT&RUN Assays"],"summary":"\n \n \n Type: Nuclease\n \n \n Mol Wgt: 43.7 kDa\n \n \n \n ","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/694/689/Screen_Shot_2020-02-12_at_11.01.55_AM__17144.1581530752.png?c=2","alt":"CUTANA™ pAG-MNase for ChIC/CUT&RUN Workflows"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/694/689/Screen_Shot_2020-02-12_at_11.01.55_AM__17144.1581530752.png?c=2","alt":"CUTANA™ pAG-MNase for ChIC/CUT&RUN Workflows"}],"date_added":"12th Aug 2019","pre_order":false,"show_cart_action":true,"has_options":true,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":[{"id":1174,"name":"Internal Comment","value":"Excess in bottom of Venom"},{"id":1175,"name":"Internal Comment","value":"Bulk in Psylocke"}],"num_reviews":null,"weight":{"formatted":"0.01 LBS","value":0.01},"demo":false,"price":{"without_tax":{"currency":"USD","formatted":"$335.00","value":335},"price_range":{"min":{"without_tax":{"currency":"USD","formatted":"$335.00","value":335},"tax_label":"Sales Tax"},"max":{"without_tax":{"currency":"USD","formatted":"$1,295.00","value":1295},"tax_label":"Sales Tax"}},"tax_label":"Sales Tax"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=694"},{"id":883,"sku":"19-1002","name":"SNAP-CUTANA™ K-MetStat Panel","url":"https://www.epicypher.com/products/nucleosomes/snap-cutana-k-metstat-panel","availability":"","rating":null,"brand":{"name":null},"category":["Nucleosomes","Nucleosomes/SNAP-CUTANA™ Spike-in Controls","Epigenetics Kits and Reagents/CUTANA™ ChIC / CUT&RUN Assays","Epigenetics Kits and Reagents/CUTANA™ CUT&Tag Assays"],"summary":"\n \n \n \n ","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/883/911/CUTANAspike-inthumbnail_RGB_white_w_border__29403.1634679576.png?c=2","alt":"SNAP-CUTANA™ K-MetStat Panel"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/883/911/CUTANAspike-inthumbnail_RGB_white_w_border__29403.1634679576.png?c=2","alt":"SNAP-CUTANA™ K-MetStat Panel"}],"date_added":"19th Oct 2021","pre_order":false,"show_cart_action":true,"has_options":false,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":[{"id":1008,"name":"Pack Size","value":"50 Reactions"},{"id":1009,"name":"Internal Comment","value":"Excess in bottom shelf of Venom."}],"num_reviews":null,"weight":{"formatted":"0.00 LBS","value":0},"demo":false,"add_to_cart_url":"https://www.epicypher.com/cart.php?action=add&product_id=883","price":{"without_tax":{"currency":"USD","formatted":"$295.00","value":295},"tax_label":"Sales Tax"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=883"},{"id":758,"sku":"13-0042","name":"CUTANA™ IgG Negative Control Antibody for CUT&RUN and CUT&Tag","url":"https://www.epicypher.com/products/nucleosomes/snap-cutana-spike-in-controls/cutana-igg-negative-control-antibody-for-cut-run-and-cut-tag","availability":"","rating":null,"brand":{"name":null},"category":["Nucleosomes/SNAP-CUTANA™ Spike-in Controls","Antibodies/CUTANA™ CUT&RUN Antibodies","Epigenetics Kits and Reagents","Epigenetics Kits and Reagents/CUTANA™ ChIC / CUT&RUN Assays","Epigenetics Kits and Reagents/CUTANA™ CUT&Tag Assays"],"summary":"\n \n \n Type: Polyclonal\n \n \n Host: Rabbit\n \n \n Applications: CUT&R","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/758/739/1579731877.1280.1280__61771.1592491196.png?c=2","alt":"CUTANA™ IgG Negative Control Antibody for CUT&RUN and CUT&Tag"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/758/739/1579731877.1280.1280__61771.1592491196.png?c=2","alt":"CUTANA™ IgG Negative Control Antibody for CUT&RUN and CUT&Tag"}],"date_added":"17th Jun 2020","pre_order":false,"show_cart_action":true,"has_options":false,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":[{"id":1148,"name":"Pack Size","value":"100 µg"},{"id":1149,"name":"Internal Comment","value":"bulk stocks from Thermo in cold room"}],"num_reviews":null,"weight":{"formatted":"0.01 LBS","value":0.01},"demo":false,"add_to_cart_url":"https://www.epicypher.com/cart.php?action=add&product_id=758","price":{"without_tax":{"currency":"USD","formatted":"$105.00","value":105},"tax_label":"Sales Tax"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=758"}],"shipping_messages":[],"rating":0,"meta_keywords":"H3K4me3 antibody, histone H3K4me3 antibody, histone H3 lysine 4 trimethylated antibody, CUT&RUN antibody, CUT&Tag antibody","show_quantity_input":1,"title":"H3K4me3 Antibody, SNAP-Certified™ for CUT&RUN and CUT&Tag","gift_wrapping_available":false,"min_purchase_quantity":0,"customizations":[],"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/1119/1196/snap-chip-ab__70507.1557259520.1280.1280__90586.1575483123.1280.1280__88802__68533.1701245530.png?c=2","alt":"H3K4me3 Antibody, SNAP-Certified™ for CUT&RUN and CUT&Tag"}]} Pack Size: 100 µg
- Type: Monoclonal [2909-3D7]
- Host: Rabbit
- Applications: CUT&RUN, CUT&Tag
- Reactivity: Human, Wide Range (Predicted)
- Format: Protein A affinity-purified
- Target Size: 15 kDa
Description
This H3K4me3 (histone H3 trimethylated at lysine 4) antibody meets EpiCypher’s lot-specific SNAP-Certified™ criteria for specificity and efficient target enrichment in both CUT&RUN and CUT&Tag applications. This requires <20% cross-reactivity to related histone PTMs determined using the SNAP-CUTANA™ K-MetStat Panel of spike-in controls (EpiCypher 19-1002, Figures 1 and 5). High target efficiency is confirmed by consistent genomic enrichment at varying cell inputs: 500k and 50k cells in CUT&RUN (Figures 2-3); 100k and 10k cells in CUT&Tag (Figures 6-7). High efficiency antibodies display similar peak structures (Figures 3 and 7) and highly conserved genome-wide signal (Figures 2 and 6) even at reduced cell numbers. This antibody targets histone H3K4me3, which is enriched at active promoters near transcription start sites (TSS) and promotes gene activation.
Validation Data - CUT&RUN
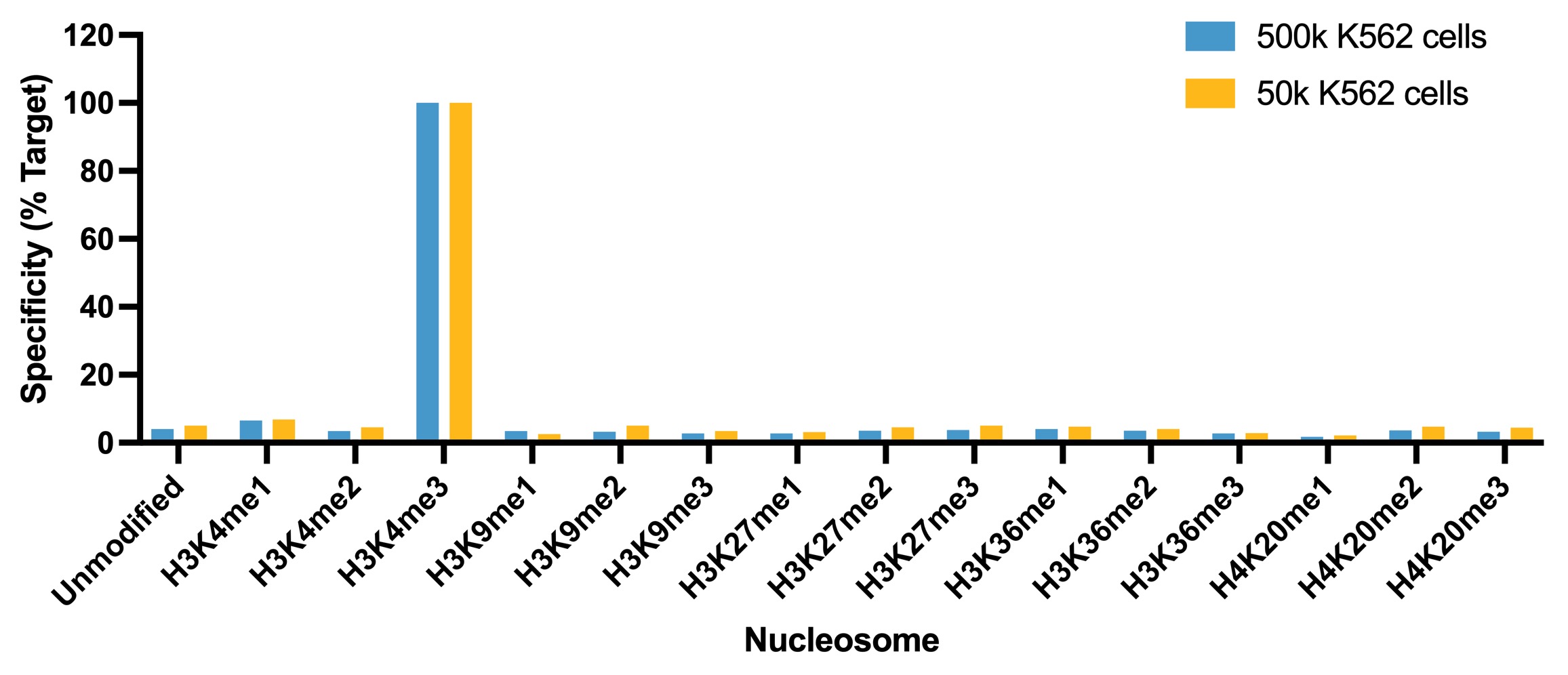
Figure 1: SNAP specificity analysis in
CUT&RUN
CUT&RUN was performed as described in
Figure 4. CUT&RUN sequencing reads
were aligned to the unique DNA barcodes corresponding
to each nucleosome in the K-MetStat panel (x-axis).
Data are expressed as a percent relative to on-target
recovery (H3K4me3 set to 100%). The antibody showed highly specific recovery of H3K4me3 spike-in
nucleosomes at both 500k and 50k cells.
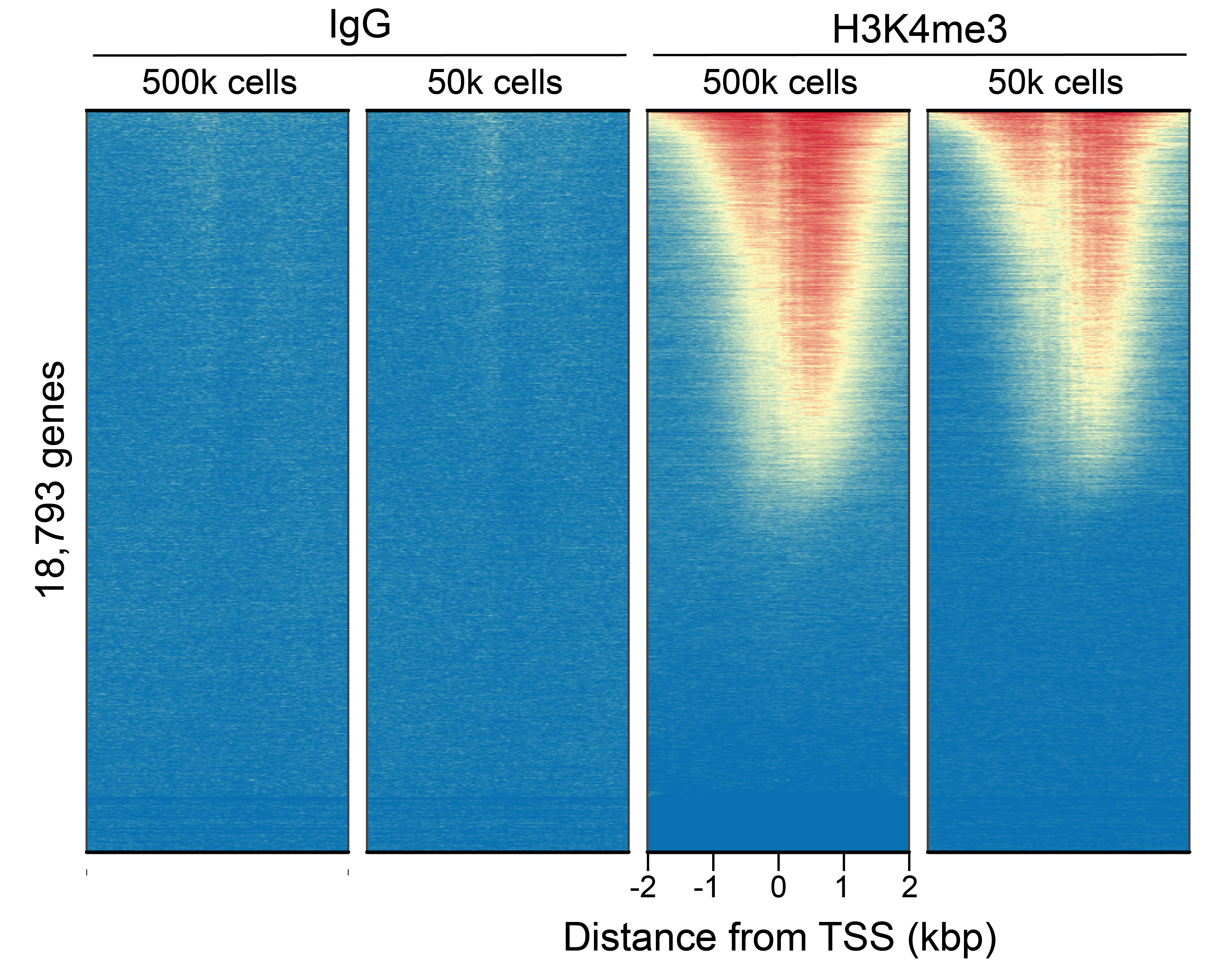
Figure 2: CUT&RUN genome-wide
enrichment
CUT&RUN was performed as described in
Figure 4. Sequence reads were aligned
to 18,793 annotated transcription start sites (TSSs ±
2 kbp). Signal enrichment was sorted from highest to
lowest (top to bottom) relative to the H3K4me3 - 500k
cells reaction (all gene rows aligned). High, medium,
and low intensity are shown in red, yellow, and blue,
respectively. H3K4me3 antibodies produced the expected
enrichment pattern, which was consistent between 500k
and 50k cells and greater than the IgG negative
control.
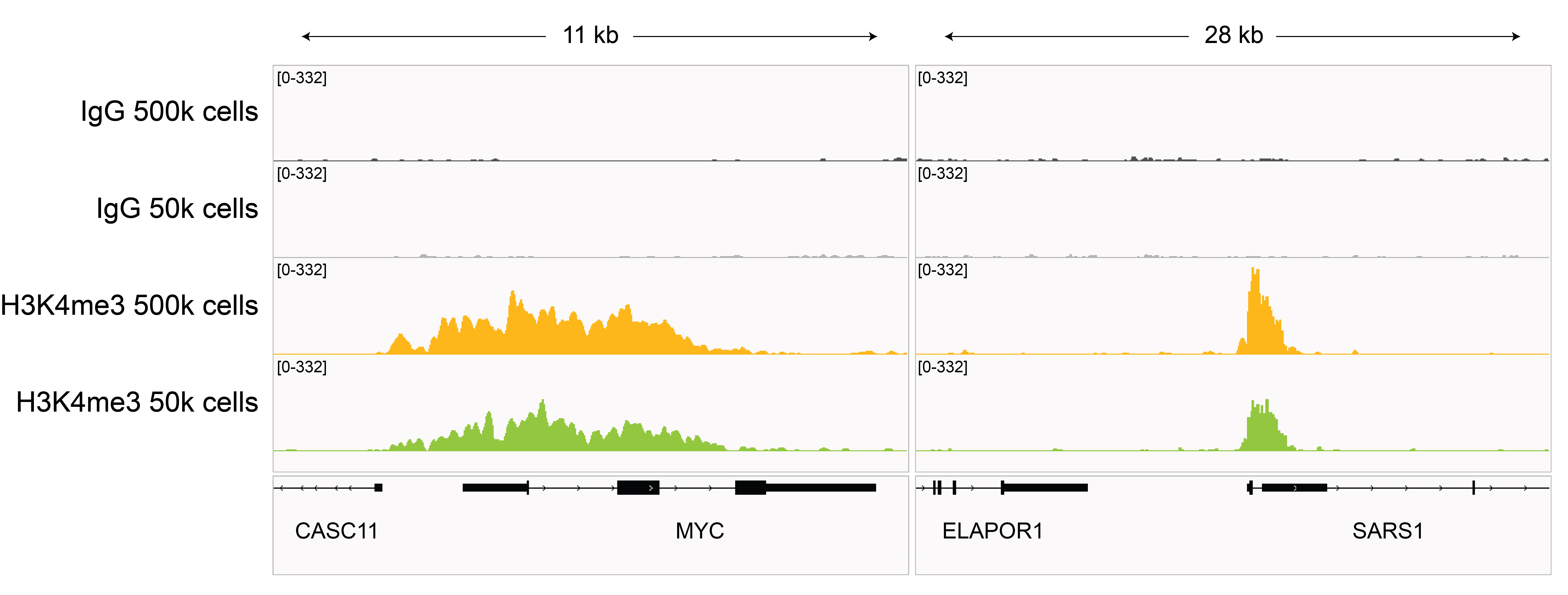
Figure 3: H3K4me3 CUT&RUN representative browser tracks
CUT&RUN was performed as described in
Figure 4. Gene browser shots were
generated using the Integrative Genomics Viewer (IGV,
Broad Institute). H3K4me3 antibody tracks display sharp peaks at gene promoters, consistent with the
biological function of this PTM. Similar results in peak structure and
location were observed for both 500k and 50k cell
inputs.
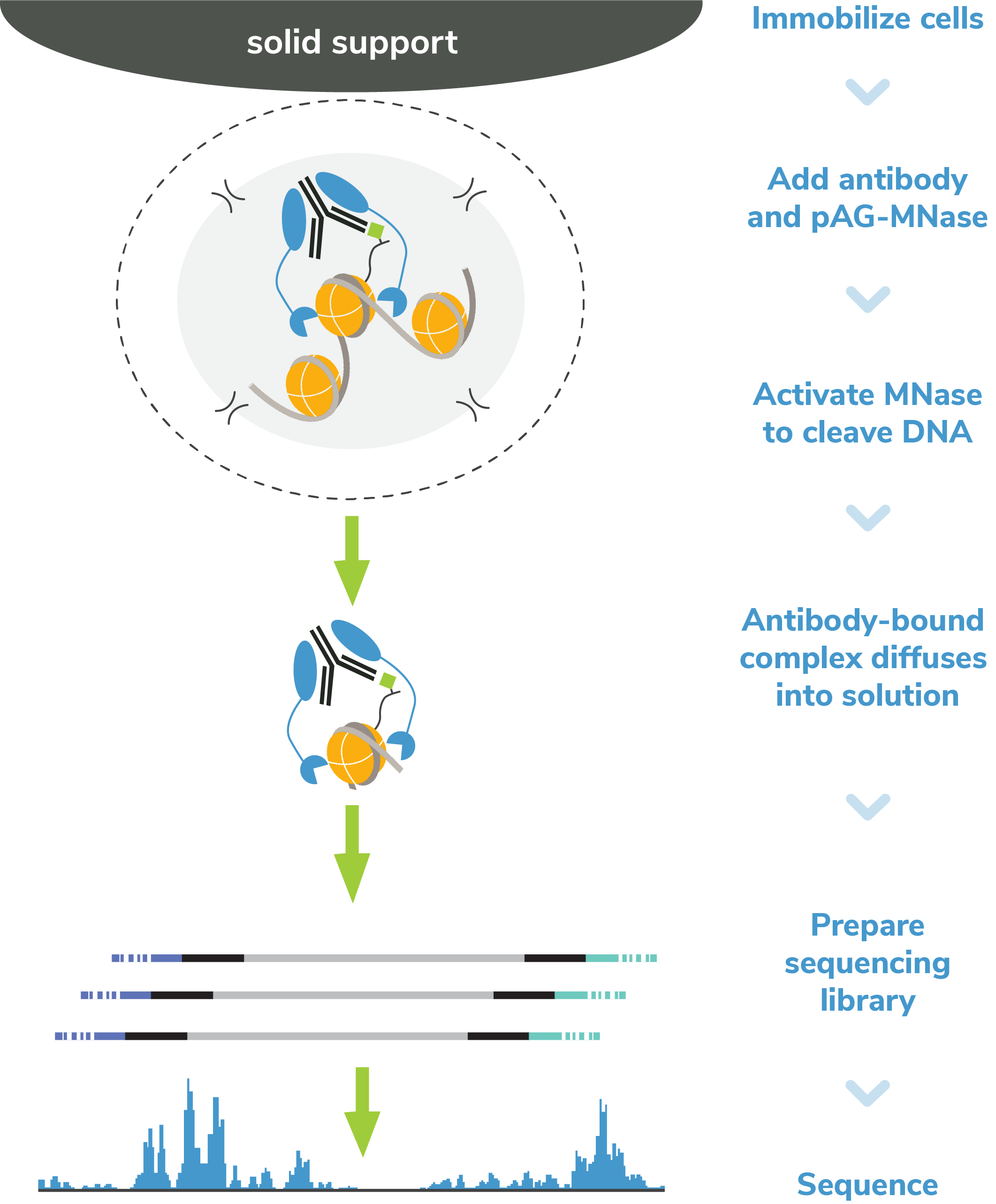
Figure 4: CUT&RUN methods
CUT&RUN was performed on 500k and 50k K562 cells with
the SNAP-CUTANA™ K-MetStat Panel (EpiCypher
19-1002) spiked-in prior to the addition of 0.5 µg of either
IgG negative control (EpiCypher
13-0042) or H3K4me3 antibodies. The experiment was performed
using the CUTANA™ ChIC/CUT&RUN Kit v3 (EpiCypher
14-1048). Library preparation was performed with 5 ng of
CUT&RUN enriched DNA (or the total amount recovered if
less than 5 ng) using the CUTANA™ CUT&RUN Library Prep
Kit (EpiCypher
14-1001/14-1002). Both kit protocols were adapted for high
throughput Tecan liquid handling. Libraries were run
on an Illumina NextSeq2000 with paired-end sequencing
(2x50 bp). Sample sequencing depth was 3.5 million reads (IgG 500k cell input), 4.0 million reads
(IgG 50k cell input), 5.6 million reads (H3K4me3 500k cell input), and 2.8 million reads (H3K4me3
50k cell input). Data were
aligned to the hg19 genome using Bowtie2. Data were
filtered to remove duplicates, multi-aligned reads,
and ENCODE DAC Exclusion List regions.
Validation Data - CUT&Tag
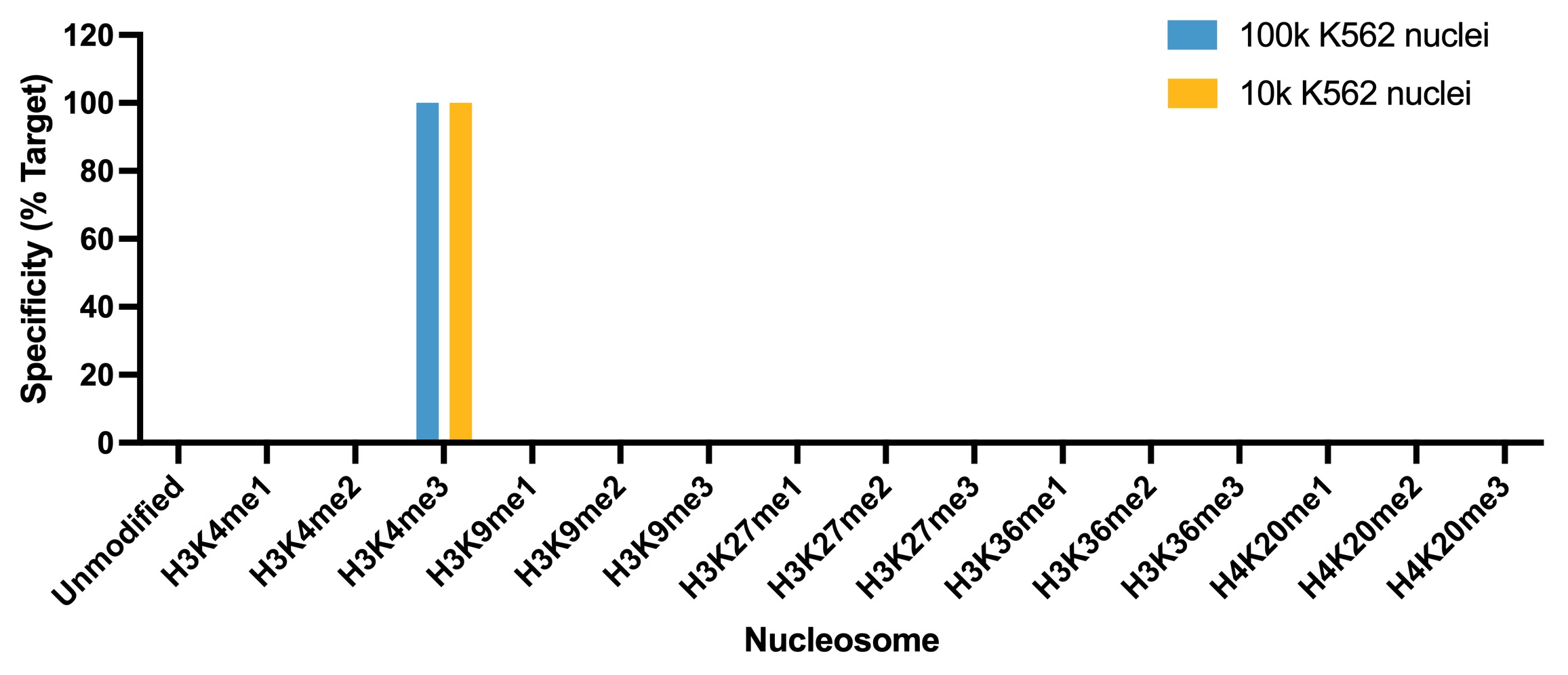
Figure 5: SNAP specificity analysis in
CUT&Tag
CUT&Tag was performed as described in
Figure 8. CUT&Tag sequencing reads
were aligned to the unique DNA barcodes corresponding
to each nucleosome in the K-MetStat panel (x-axis).
Data are expressed as a percent relative to on-target
recovery (H3K4me3 set to 100%). The antibody showed highly specific recovery of H3K4me3 spike-in
nucleosomes at both 100k and 10k nuclei.
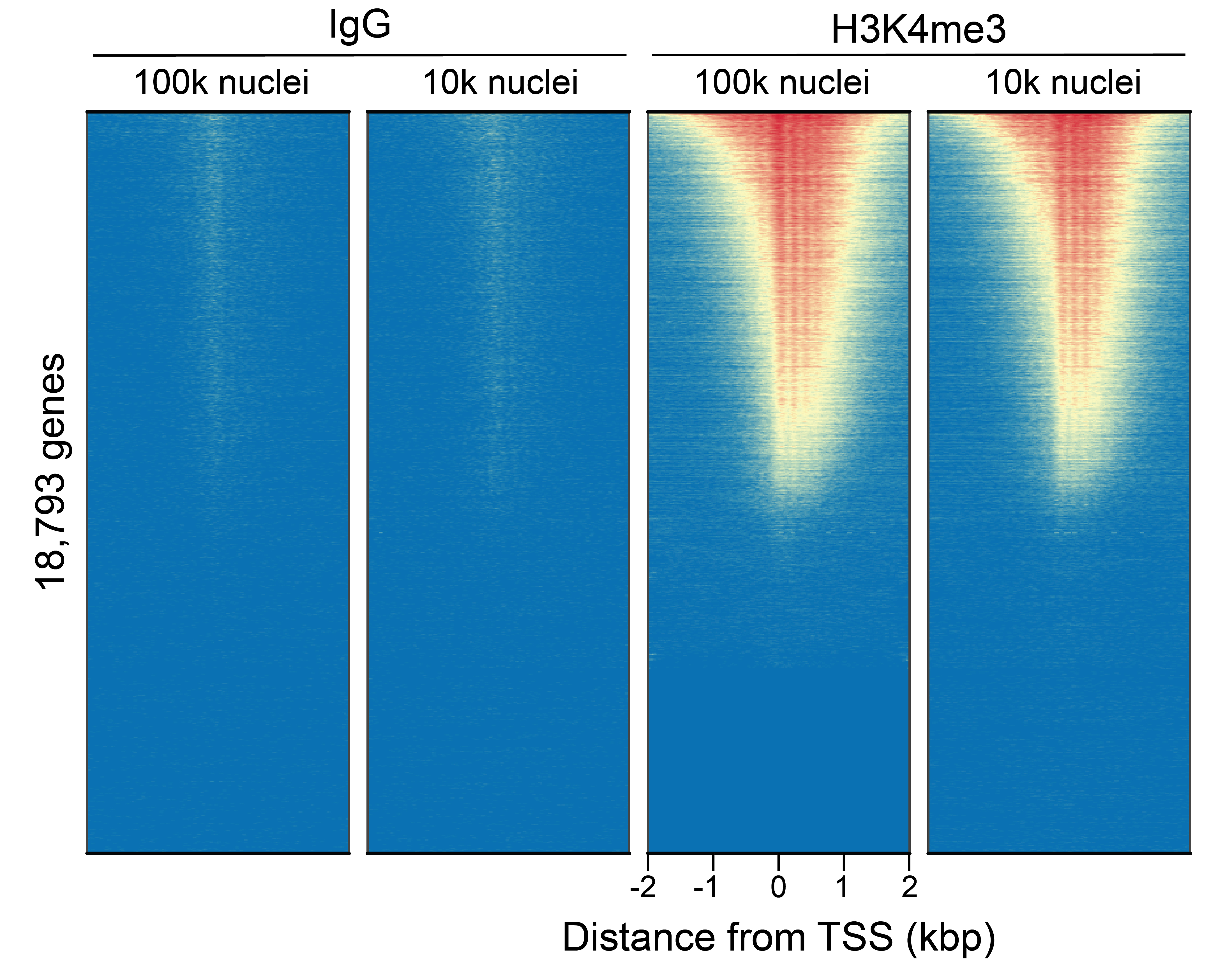
Figure 6: CUT&Tag genome-wide enrichment
CUT&Tag was performed as described in
Figure 8. Sequence reads were aligned
to 18,793 annotated transcription start sites (TSSs ±
2 kbp). Signal enrichment was sorted from highest to
lowest (top to bottom) relative to the H3K4me3 - 100k
nuclei reaction (all gene rows aligned). High, medium,
and low intensity are shown in red, yellow, and blue,
respectively. H3K4me3 antibodies produced the expected
enrichment pattern, which was consistent between 100k
and 10k nuclei and greater than the IgG negative
control.
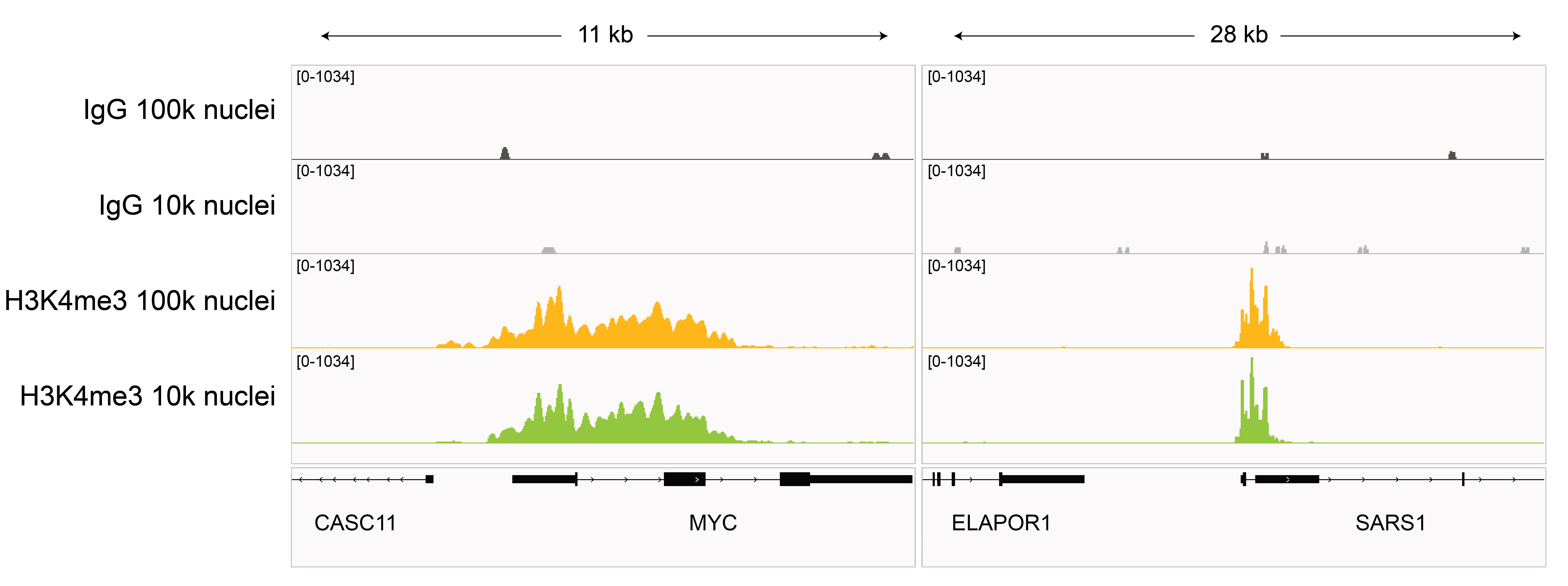
Figure 7: H3K4me3 CUT&Tag representative browser tracks
CUT&Tag was performed as described in
Figure 8. Gene browser shots were
generated using the Integrative Genomics Viewer (IGV,
Broad Institute). H3K4me3 antibody tracks display sharp peaks at gene promoters, consistent with the
biological function of this PTM. Similar results in peak structure and
location were observed for both 100k and 10k nuclei
inputs.
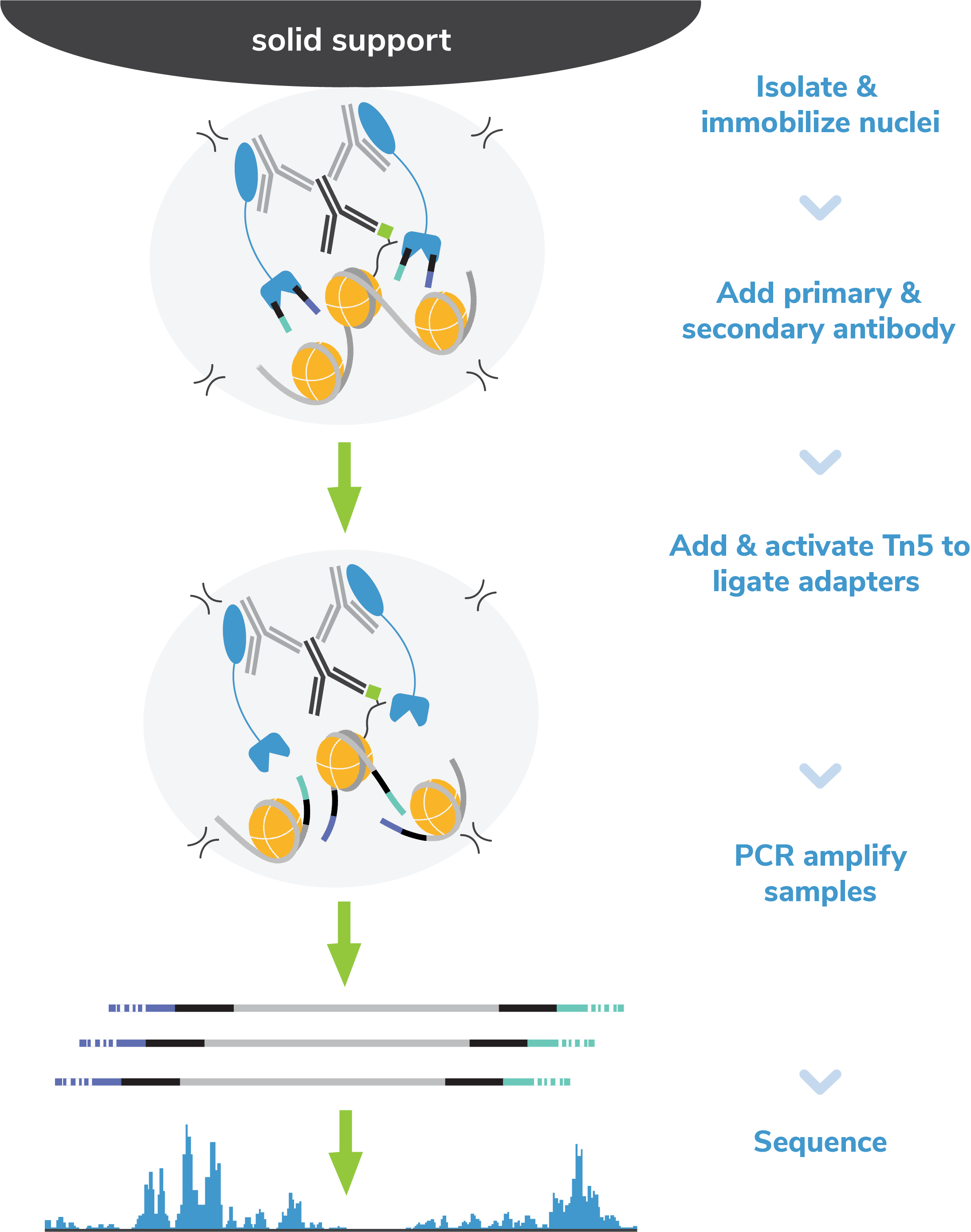
Figure 8: CUT&Tag methods
CUT&Tag was performed on 100k and 10k K562 nuclei with
the SNAP-CUTANA™ K-MetStat Panel (EpiCypher
19-1002) spiked-in prior to the addition of 0.5 µg of either
IgG negative control (EpiCypher
13-0042) or H3K4me3 antibodies. The experiment was performed
using the CUTANA™ CUT&Tag Kit v1 (EpiCypher
14-1102/14-1103). Libraries were run on an Illumina NextSeq2000 with
paired-end sequencing (2x50 bp). Sample sequencing depth was 1.3 million reads (IgG 100k nuclei
input), 2.0 million reads (IgG 10k nuclei input), 6.9 million reads (H3K4me3 100k nuclei input), and
10.6 million reads (H3K4me3 10k nuclei input). Data were aligned to the
hg19 genome using Bowtie2. Data were filtered to
remove duplicates, multi-aligned reads, and ENCODE DAC
Exclusion List regions.